406.587.4531
simmental@simmgene.com
406.587.4531
simmental@simmgene.com
Simple Trait Selection
Coat Color and Coat Color Dilution
- All animals have two copies of each gene, and pass along one copy to their offspring.
- An individual with the same two alleles for a gene is homozygous. An individual with two different alleles of a gene is heterozygous.
- In traits with complete dominance, like coat color, an animal needs only one of the dominant alleles to display the dominant phenotype. An animal needs two recessive alleles to display the recessive phenotype.
- Some of the cells within the Punnett square will have different genotypes but the same phenotype. For instance, with a completely dominant trait, the heterozygotes will appear the same as the homozygote dominant animals.
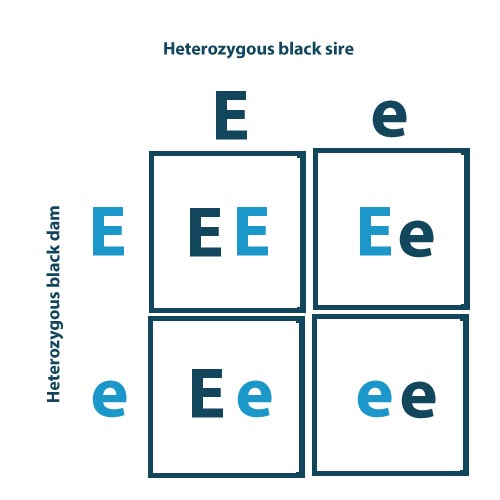
Example: Coat color, a simply inherited trait that exhibits complete dominance.
- Alleles: E = black and dominant, e = red and recessive
- Mating: Parents that are heterozygous have the same 50% chance of passing on the dominant allele as they do the recessive. So, when mating a heterozygous black sire with a heterozygous black dam, there is a 25% chance the progeny will be homozygous dominant, a 50% chance the progeny will be heterozygous, and a 25% chance that the progeny will be homozygous recessive.
Wild Type
Wild Type is a generic term in genetics referring to the normal allele or in this case the original allele. In the extension locus, the wild type variant is the original DNA sequence, and the black and red variants are mutations of the wild type sequence. The Wild Type allele (E+) can produce varying degrees of red/yellow to brown/black. The order of dominance for these alleles is thought to be E > E+ > e; in other words, black is dominant to wild type which is dominant to red. However, a new study suggests that in some cattle, particularly in Bos indicus x Bos taurus crosses, black is not completely dominant to wild type.
Jersey, Brown Swiss, Tarentaise, Texas Longhorn, Brahman, and other Zebu cattle carry the wild type allele, but it is not limited to these populations (note picture of the E+/e SimAngus bull). Due to its prevalence in the Brahman breed, Simbrah cattle frequently carry the wild type variant. As wild type animals have the ability to make both red or black hair, their coat color can be more variable. Homozygous wild type cattle range in coat color from yellow to black, although the most common coloration is reddish brown or brownish black. Frequently, wild type animals become darker as they age, and wild type bulls are typically darker-pigmented than wild type cows (see pictures for examples). Wild type animals commonly have darker pigmentation at the feet, head, and neck, and have a tan ring around the muzzle.
As wild type animals can make black or red hair, other genes that affect the ratio of pheomelanin to eumelanin production will affect wild type animals but not black or true red (ee) cattle. Cattle can be blazed, spotted, brindled, roan, brockle, belted, diluted, dun, and the list goes on and on. There are many genes involved in these variations of coat color.
The following table represents the genotype possessed by the animal and thee resulting phenotype (coat color appearance):
ED/E+ = Black
E+/E+ = Various; reddish brown, to brownish black
E+/e = Red or red/black
The wild type coat color test is included in the regular coat color DNA test offered by ASA. If you have any questions, contact the DNA Department at dna@simmgene.com.

Photo Description: SimAngus Bull (3/4 Simmental, ¼ Red Angus) that is a wild type carrier (E+/e).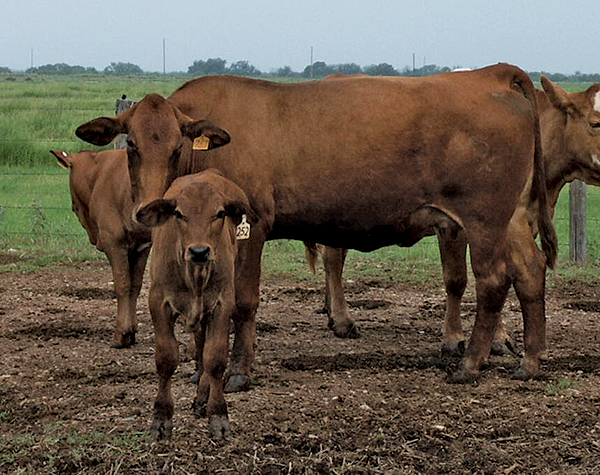 Photo Description: Simbrah cow (above) is a Wild type carrier (E+/e). Her bull calf, a likely wild type carrier, started with a similar reddish color but developed a darker pigmentation, especially on the head and neck and feet as he aged (pictured below).
Photo Description: Simbrah cow (above) is a Wild type carrier (E+/e). Her bull calf, a likely wild type carrier, started with a similar reddish color but developed a darker pigmentation, especially on the head and neck and feet as he aged (pictured below).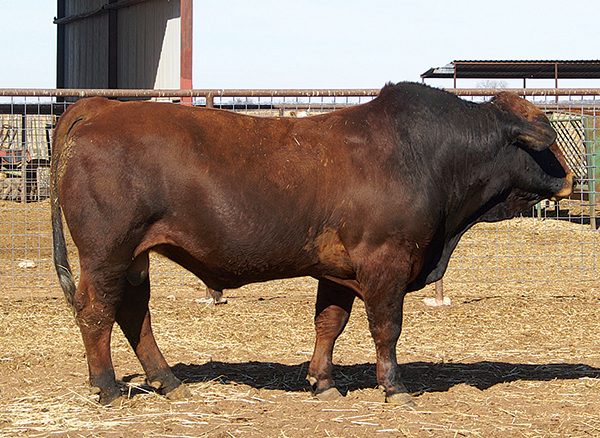
Dilution
Like the black coat color phenotype, dilution is a dominant trait, so it only takes one mutated gene to see the dilution. Let’s call the dilution mutation “D” and the normal Pmel17 allele “d.” If an animal carries one copy of the dilution mutation and one normal allele for the Pmel17 gene, you would expect that half of their progeny will inherit the dilution mutation and the other half the normal allele. Whether or not the progeny appear to have a diluted coat color is dependent on the coat color of the offspring. All the black calves with the dilution mutation will have a gray coat color. The black calves with the normal Pmel17 gene will be black. All the red calves will most likely appear red (although those with the diluter gene may have a slightly lighter red color). The following list represents the genotype (which alleles the animal has; E = black coat, e = red coat, D = dilution mutation, d = no dilution) and the resulting phenotype (coat color appearance):
Ee/dd = Black
Ee/Dd = Gray
ee/dd = Red
ee/Dd = Red (might be slightly lighter in color)
Why is it relevant to understand the above information? If you have animals that are black, you probably don’t need to test them for the diluter gene as they would appear gray if they carried it. However, your red animals can “hide” the dilution mutation without notice. Therefore, if you are mating red animals to black animals and you don’t want any gray calves, you should test your red animals for the diluter gene.
To order the dilutor test, please contact the ASA at DNA@simmgene.com.
Genetic Conditions
Known as “Curly Calf Syndrome,” AM results in stillborn calves small in size with diminished muscling, bent limbs, and twisted spines.
Recessive, lethal, affecting Angus and Angus-influenced cattle.
A genetic mutation is a change in the genetic code from what previously existed. While some genetic mutations are advantageous (polled, for example), the majority of mutations in nature tend to hinder a population’s success via harmful or lethal means. Mutations of this nature are often referred to as genetic conditions or genetic defects.
Dominant mutations always influence an animal’s phenotype so the mutation can easily be selected for or against. Recessive mutations, however, tend to exist in a population even when harmful to the point of being lethal. This is because animals can carry the recessive gene without showing any signs of it. When carrier animals are mated to other carriers, the resulting offspring have a chance of showing symptoms. Fortunately, technology has evolved to the point at which animals that appear normal yet are carriers of recessive genetic conditions can be identified if a DNA test exists for that mutation.
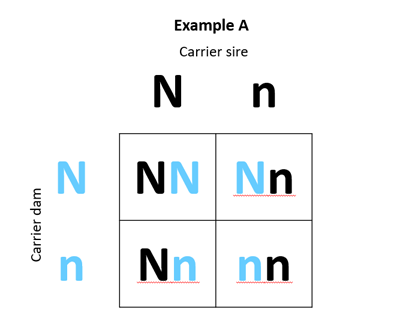
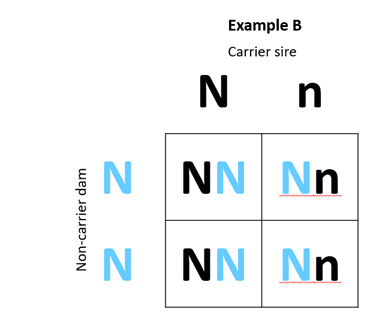
A 2 x 2 Punnett Square can be used to illustrate the outcomes of various matings. In Example A below, we have mated a carrier sire to a carrier dam, while in Example B we have mated a carrier sire to a non-carrier dam.
Horned/Polled/Scurred
Let’s use the following abbreviations for the horned, polled, scurred, and no scurs alleles:
P = polled allele and dominant
p = horned allele and recessive
S = no scurs (a different gene than horned/polled gene)
s = scurred (a different gene than horned/polled gene)
Horned/polled and scurred genotypes and the resulting phenotypes:
All homozygous polled (PP) animals are phenotypically polled independent of the scurred gene.
All homozygous horned (pp) animals have horns and are unaffected by the scurred gene.
The heterozygous animals at the horned/polled gene (Pp) can either be polled or have scurs depending on their sex and the alleles for the scurred gene.
Scurred Genotype | Cow Phenotype | Bull Phenotype |
|---|---|---|
SS | Polled | Polled |
Ss | Polled | Scurred |
ss | Scurred | Scurred |
*All animals in this table are heterozygous for the horned/polled (Pp) gene. See the above text for more explanation.
At present, there is no test for the scurred gene but you can test your cattle for the Horned/polled alleles through the ASA. To order tests send inquiries to DNA@simmgene.com.
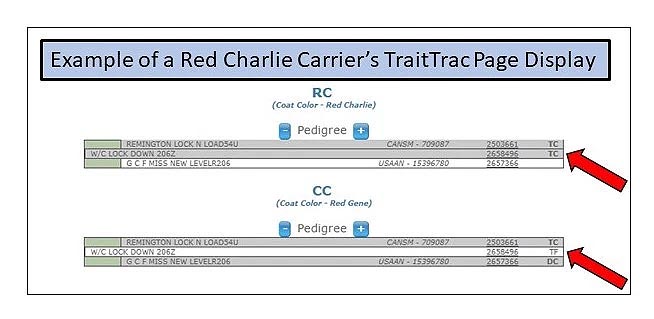
Red Charlie: A Newly Discovered Red Coat Color Variant
Subscribe to eNews ASA's weekly newsletter.
Sales Call: Calendar of upcoming sales and
eBlast: Sales and other industry promotions